SCIP: scalable cytometry image processing using Dask in a high performance computing environment
A software for distributed processing of bioimaging datasets
- Track: HPC, Big Data, and Data Science devroom
- Room: D.hpc
- Day: Sunday
- Start: 10:30
- End: 11:00
- Video with Q&A: D.hpc
- Video only: D.hpc
- Chat: Join the conversation!
Bioimage analysis is the process of extracting novel insights from microscopy images of tissues, cells or other biological entities. Many tools, such as ImageJ, QuPath or CellProfiler are heavily used by researchers to quantitatively interpret those complex images. These tools perform tasks such as normalization, image segmentation, image masking or feature extraction. However, these tools are designed for usage on local workstations with a GUI and for the most part only allow vertical scaling to deal with an increase in dataset size. This limited scalability poses a problem as bioimaging datasets keep growing in volume.
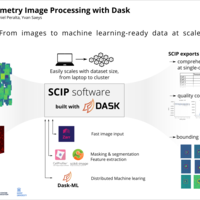
Here, we introduce Scalable Cytometry Image Processing (SCIP). SCIP is an open-source tool that implements single-cell bioimage processing on top of Dask, a framework for distributed computing written in Python. By utilizing Dask to execute all computations, scalability is integral to the software allowing it to be executed on high performance computing clusters. Dask's smart task scheduling ensures computational resources are used efficiently. SCIP also takes advantage of Dask features such as fault tolerance, load balancing or data locality. This allows SCIP to process large datasets more efficiently and more robustly compared to other tools.
SCIP was written in Python, and all code is freely available. It can run in local (single-node) or distributed mode. In the latter, Dask's components communicate efficiently using the MPI standard. SCIP can be used as a stand-alone command line tool, or integrated into existing Python scripts using the API.
For a similar setup, SCIP showed a 4-fold decrease in runtime compared to CellProfiler. SCIP needed considerably less manual work to prepare the dataset for processing thanks to more efficient data input. Comparisons were executed on the Flemish Supercomputer Center Tier-1 high performance cluster. We processed two cytometry datasets containing images of human blood cells: an imaging flow cytometry dataset containing 270,000 6-channel images of 90 by 90 pixels, and a confocal microscopy dataset containing 869 5-channel images of 1600 by 900 pixels.
The imaging flow cytometry dataset is processed by SCIP in 484 seconds using 80 workers running on 3 compute nodes. Each node has 24 cores (Intel Xeon E5-2680v4 @ 2.4GHz) and 120GB of memory, which is divided over the workers.
The microscopy dataset is processed by SCIP in 2187 seconds using 16 workers running on 1 compute node. This node has 32 cores (Intel Xeon Silver 4110 CPU @ 2.10GHz) and 364GB memory, which is divided over the workers. This workflow also uses a GPU (Nvidia GeForce RTX 3090) for identifying the cells in the microscopy images.
Speakers
| Maxim Lippeveld |